Mitä tietoja FinnGenissä on?
Tältä sivulta löydät lisätietoja FinnGen-tutkimuskonsortioon kuuluvien tutkijoiden hyödynnettävissä olevasta yksilötason datasta englanniksi.
Longitudinal data file
The "Detailed longitudinal data file" is the main registry data file in FinnGen. It contains health register codes from six Finnish health registers and is used as an input file for the Endpointter to determine which individuals fall into each FinnGen endpoint. Endpointter is the software that the FinnGen register team uses to assign individuals to particular phenotypic endpoints.
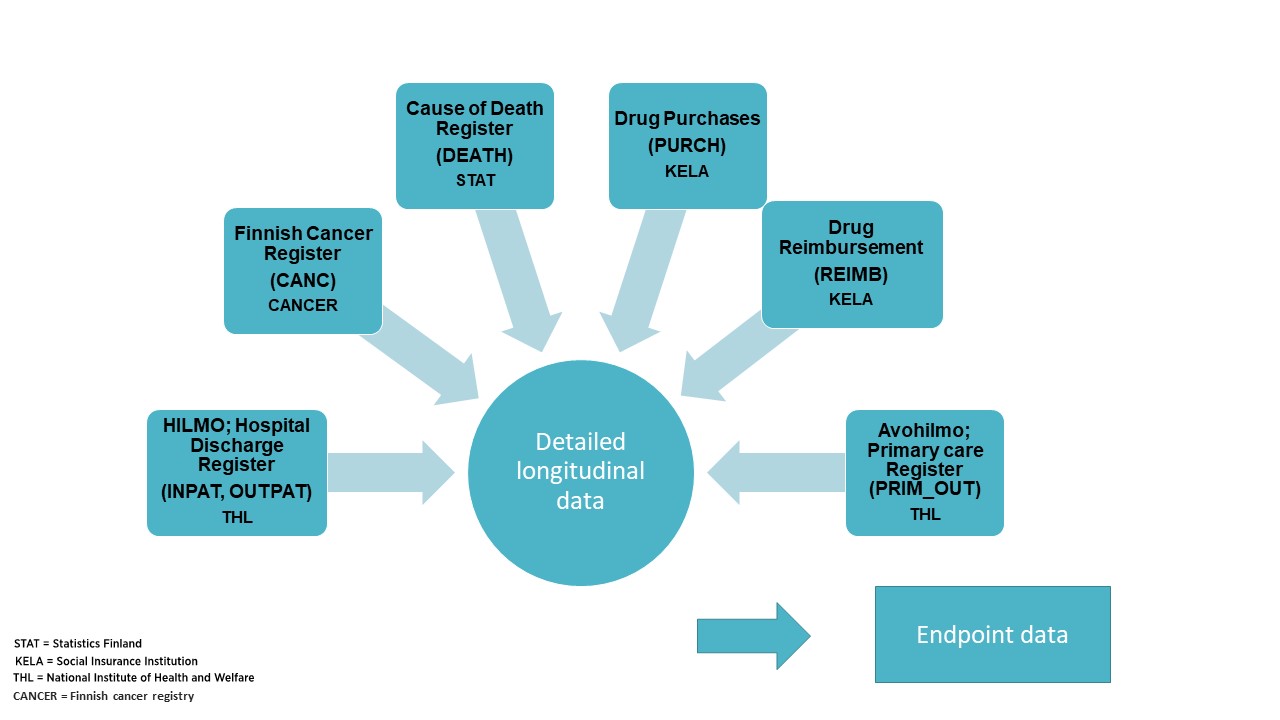
"Longitudinal" format means that the data contains several events/entries of the register codes from the same individual received at different times.
Register data from each registry is available from the beginning of the registry until end of the follow-up. Figure below presents register follow-up years, and main codes within each registry, for data freeze 6. The same pattern continues through later years as we receive more follow-up years for each register.
If you would like to learn more about the Finnish health registers, please visit the FinRegistry project's website.
Code sets in detailed longitudinal data:
Kela (Social insurance institution)
The REIMB/KELA codes are unique to Finland - these are longterm reimbursement codes for a class of drugs and imply a robust review of diagnosis.
- ATC – drug identifiers
- REIMB – drug reimbursements
- VNRO – drug product number including dosage and number in package
Hilmo/Hospital codes
- Diagnosis in ICD-10, ICD-9 and ICD-8
- Procedures (Nomesco, Finnish Hospital league*, Heart patient**)
Cancer registry
- ICD-O-3 (topology, morphology, behavior codes)
Avohilmo
- Diagnosis in ICD-10 and nurse codes ICPC2
- Procedures (SPAT, Nomesco)
- Dental codes
*) Finnish specific procedure codes, **) Finnish specific demanding heart patient procedure code
Endpoint and endpoint longitudinal data
The full catalog of the FinnGen endpoints has been created to comprehensively cover the spectrum of medical diagnoses from all medical areas. The FinnGen endpoints follow the treelike subtyping system of the ICD-10 classification system, where classification starts from the anatomy-based upper-level chapters and then divides step-by-step to more detailed subtypes.
In FinnGen, this treelike subtyping was followed to a level of detail that was seen as reasonable in the context of GWAS and PheWAS. The endpoints are named so that a prefix combining the first character of the ICD-10 diagnosis code is followed by the number of the ICD-10 chapter and then an endpoint name that aims to be as short as possible and explanatory enough to enable to reason what the endpoint is.
For example, Parkinson's disease:
- from 6th ICD-10 chapter Diseases of the nervous system
- ICD-10 code G20
-> named G6_PARKINSON
In addition to the endpoints coming quite directly from the ICD-classification, we also created endpoints according to specific requests (specific definitions according to for example age of onset, combinatory endpoints covering entities with subtypes from different chapters of the ICD-system like autoimmune diseases or alcohol-related disorders). The final sets of medical specialty specific endpoint definitions were approved by the FinnGen clinical expert groups of leading experts in their respective medical fields.
Other registry data files (additional registries)
- Vaccination registry from primary health care (vaccination, type of vaccination, disease, method and site, approx. date, contains also COVID vaccinations)
- Kidney registry (only dialysis/transplantation patients, set of diagnosis codes)
- Digital and population data services agency, birth registry (reproductive history data - detailed information on births in FinnGen participants)
- Visual impairment registry (data on legally blind patients: diagnosis, visual acuity, diameter of visual field, homonyme hemianopsia)
- Cancer screening registries (cervical and breast)
- Parental causes of death (ICD-8/9/10)
- Infectious diseases (COVID positive patients and their treatment)
- Congenital malformations (congenital chromosomal and structural anomalies, congenital hypothyroidism, teratomas)
- Service sector data (service sector (e.g. elderly home), contact type, specialty, hospital type, urgency, professional type, Kela drug reimbursement costs, Kela drug reimbursement category)
Minimum phenotype file and cohort data
Minimum phenotype file includes 15 variables. Biobanks provide age, sex, height, weight, smoking and DNA sample collection date variables. In addition, information about the region of birth, moving abroad and the number of children from the Digital and population data services agency is included. Cohort data includes information about the biobank that has delivered the sample in question to FinnGen.
Genetic data
GWAS array genotypes (500,000)
- Most of the samples have been genotyped with ThermoFisher Axiom custom array v2
- Total of 664,510 markers including:
- About 500,000 core GWAS markers
- 116,000 coding variants enriched in Finland
- >10,000 specific markers for the HLA/KIR region
- about 15,000 ClinVar variants
- about 4,600 pharmacogenomic variants
- 57,000 selected markers of special interest for partners
- 80-90,000 samples genotypes with various other GWAS arrays (Legacy genotypes)
Imputed genotypes (500,000)
- ~21,3M variants per individual
- imputed using Finnish specific WGS reference panel of ~9000 individuals